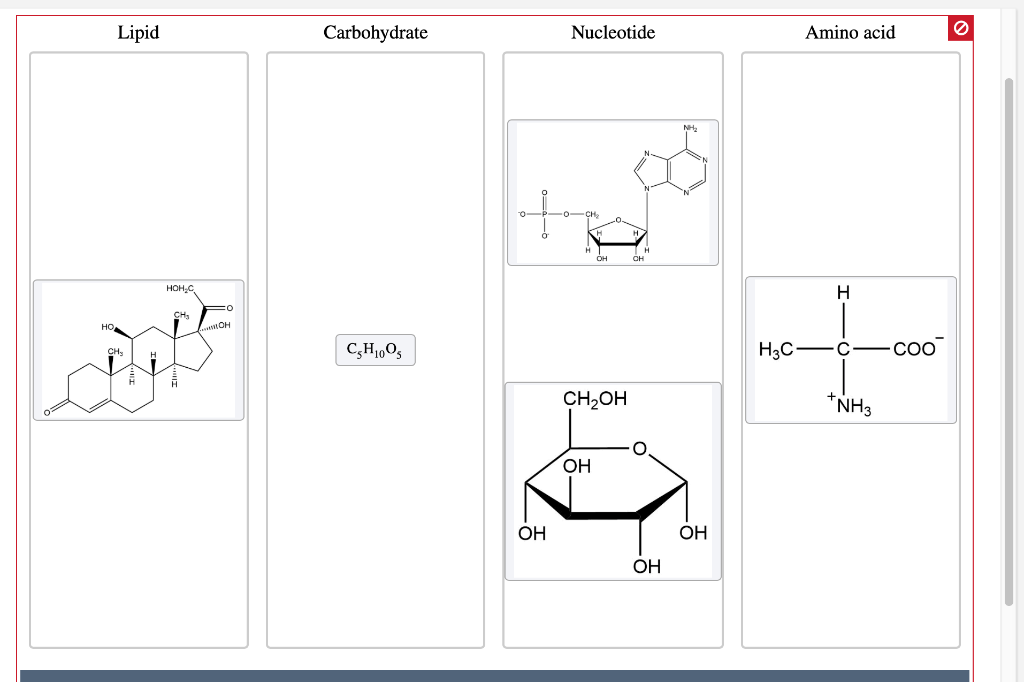
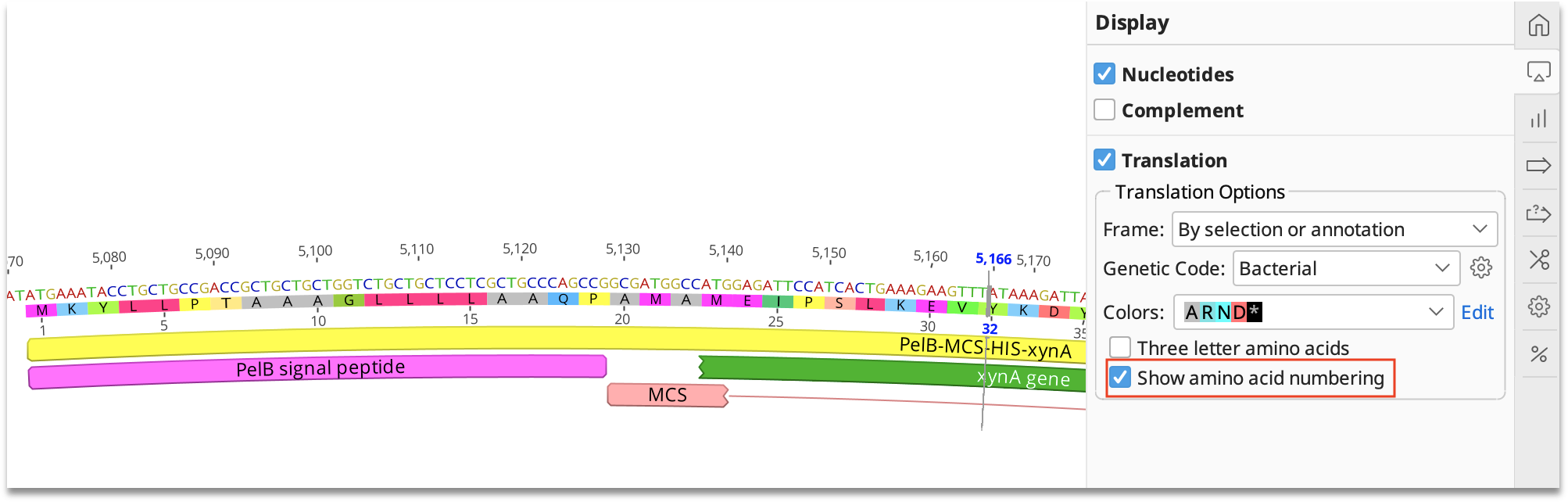
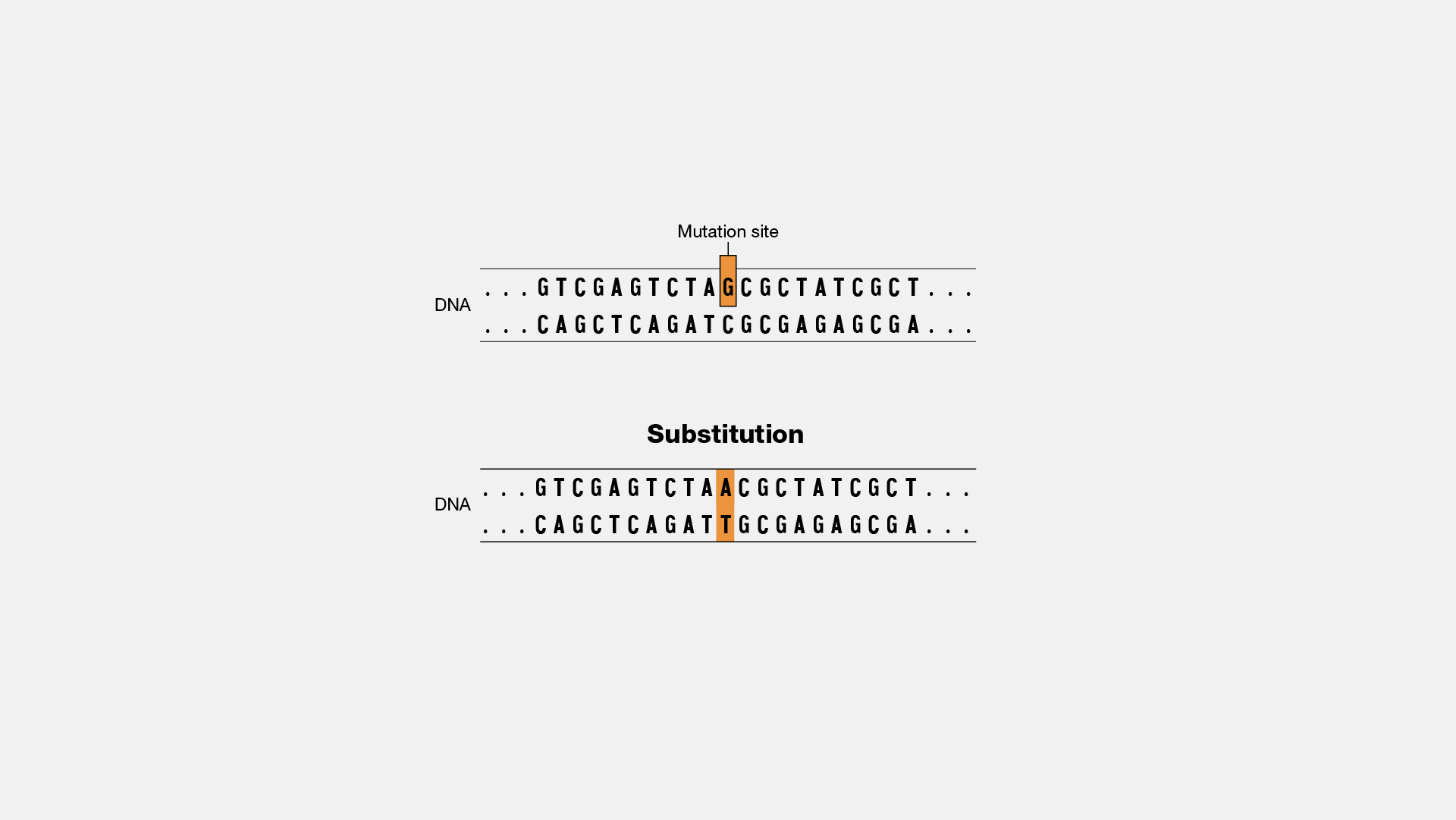
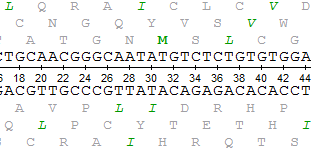
Nucleotide sequences and corresponding amino acid sequences around the deletions in two ccha1 (A) and two ccha2 (B) mutants.

Analyzing Amino Acid Protein Sequences and DNA Nucleotide Sequences to Support Evolution Practice | Biology Practice Problems | Study.com
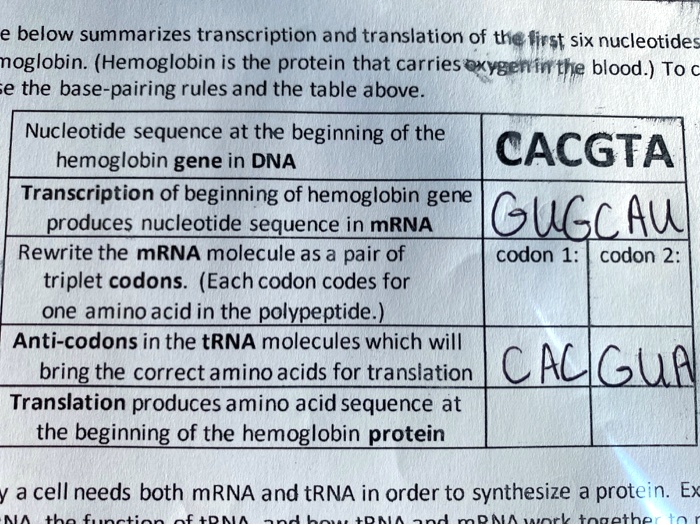

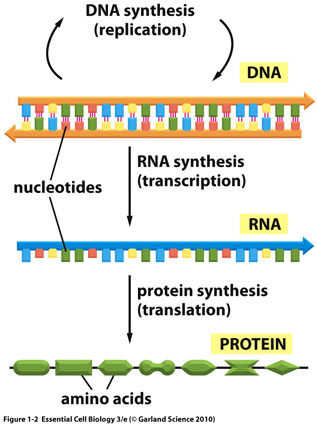
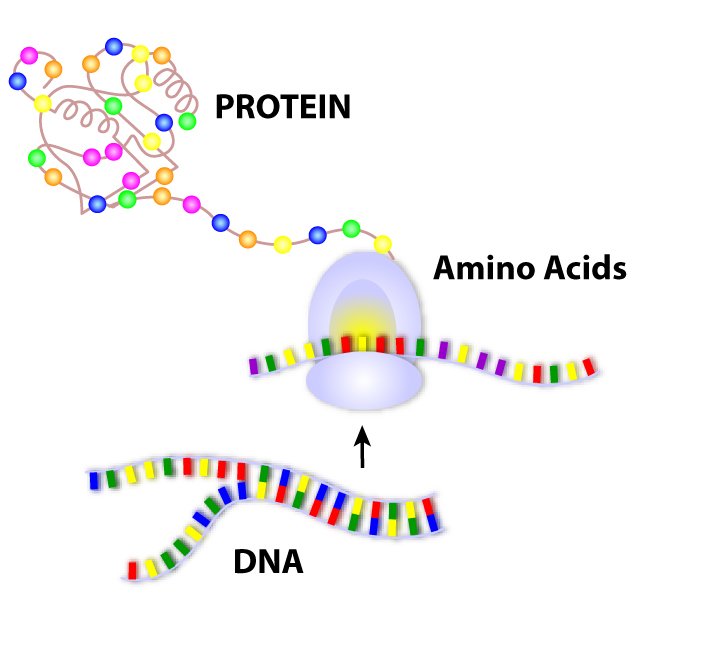
SOLVED: The text below summarizes the transcription and translation of the first six nucleotides of hemoglobin: (Hemoglobin is the protein that carries oxygen in the blood). To understand the base-pairing rules and

The given table shows the genetic code depicting the amino acids that correspond to mRNA codons. Each codon is read from 3^' (first nucleotide) to 5^' (third nucleotide). Find out the amino

dna - Why do three nucleotides code for one amino acid? Why not 5 nucleotides? - Biology Stack Exchange
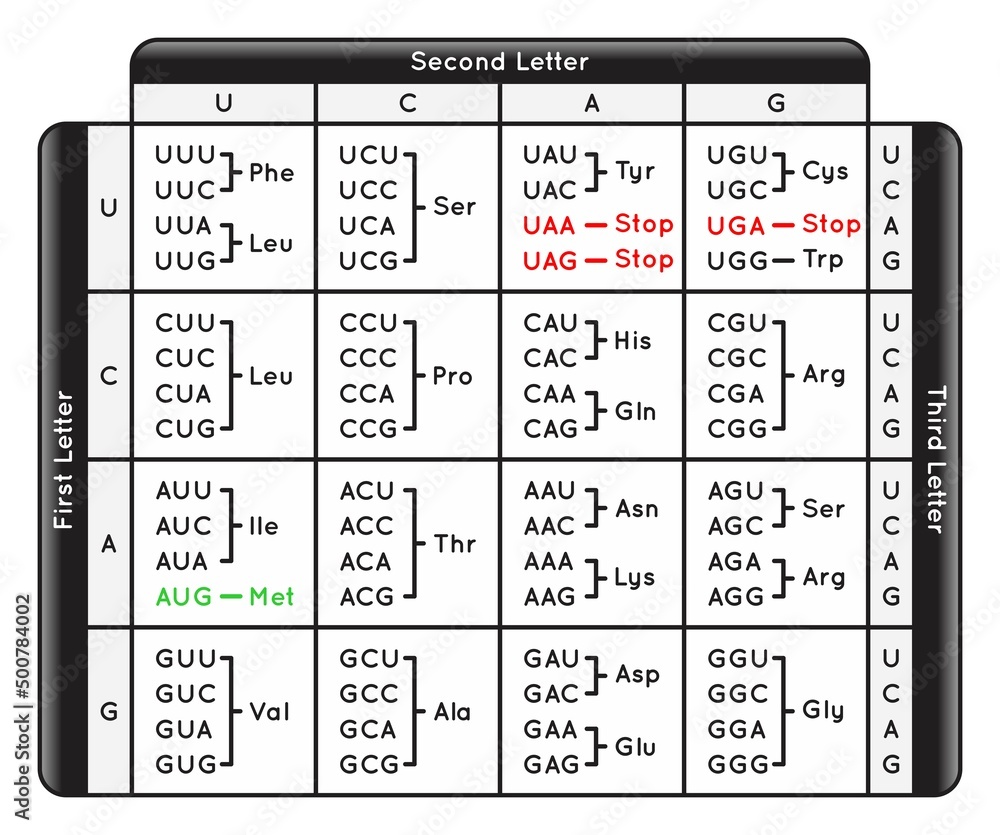
Table of Codons the Genetic Code of Human Infographic Diagram nucleotide base sequence on DNA mRNA transcription translation to protein amino acids synthesize biology omics science education vector Stock Vector | Adobe

A tri-nucleotide mapping scheme based on residual volume of amino acids for short length exon prediction using sliding window DFT method | SpringerLink

NucAmino: a nucleotide to amino acid alignment optimized for virus gene sequences | BMC Bioinformatics | Full Text
![PDF] TranslatorX: multiple alignment of nucleotide sequences guided by amino acid translations | Semantic Scholar PDF] TranslatorX: multiple alignment of nucleotide sequences guided by amino acid translations | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/d9f8f50f0010383c63e06ba8290ec60d9e37b0c7/2-Figure1-1.png)